Repository Infrastructure for Queries
Workflow Structure
Use-case Scenario: Metadata Search
Use-case Scenario: Sequence Search
Repository Infrastructure for Queries
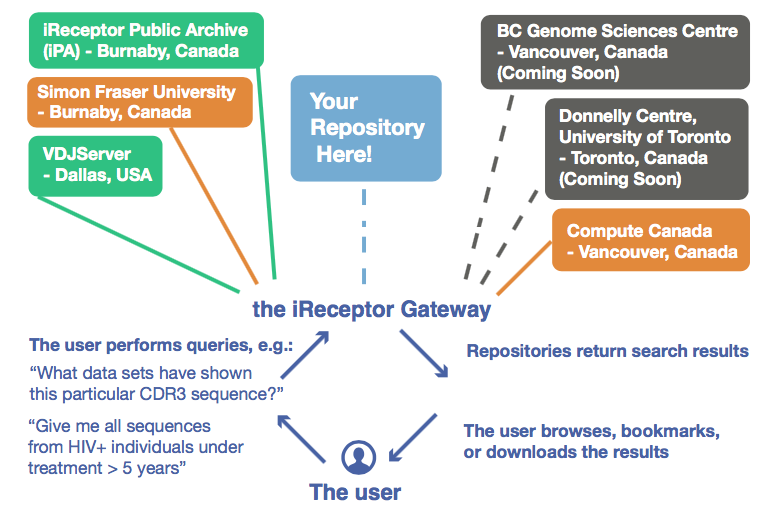
iReceptor has a federated database approach: each scientist who uploads data here holds their data sets at their own institution, under their own data and IP agreements. iReceptor users can choose how much of their data is viewable by others. Read more on iReceptor's Repository infrastructure.
Workflow Structure
iReceptor has two primary workflows: the Metadata Search and the Sequence Search. The Metadata Search workflow is for research questions about sequences' metadata, e.g. what type of study they are from, what type of cell they are from, if the subject they were taken from was healthy or not. The Sequence Search Workflow is for research questions about sequences themselves, e.g. Junction AA length or V-gene alleles. Here are two use case scenarios where different research questions necessitate different workflows:
Use-case Scenario: Metadata Search
In this scenario, a researcher would like to know: what differences are observed in T cell repertoires between child siblings? 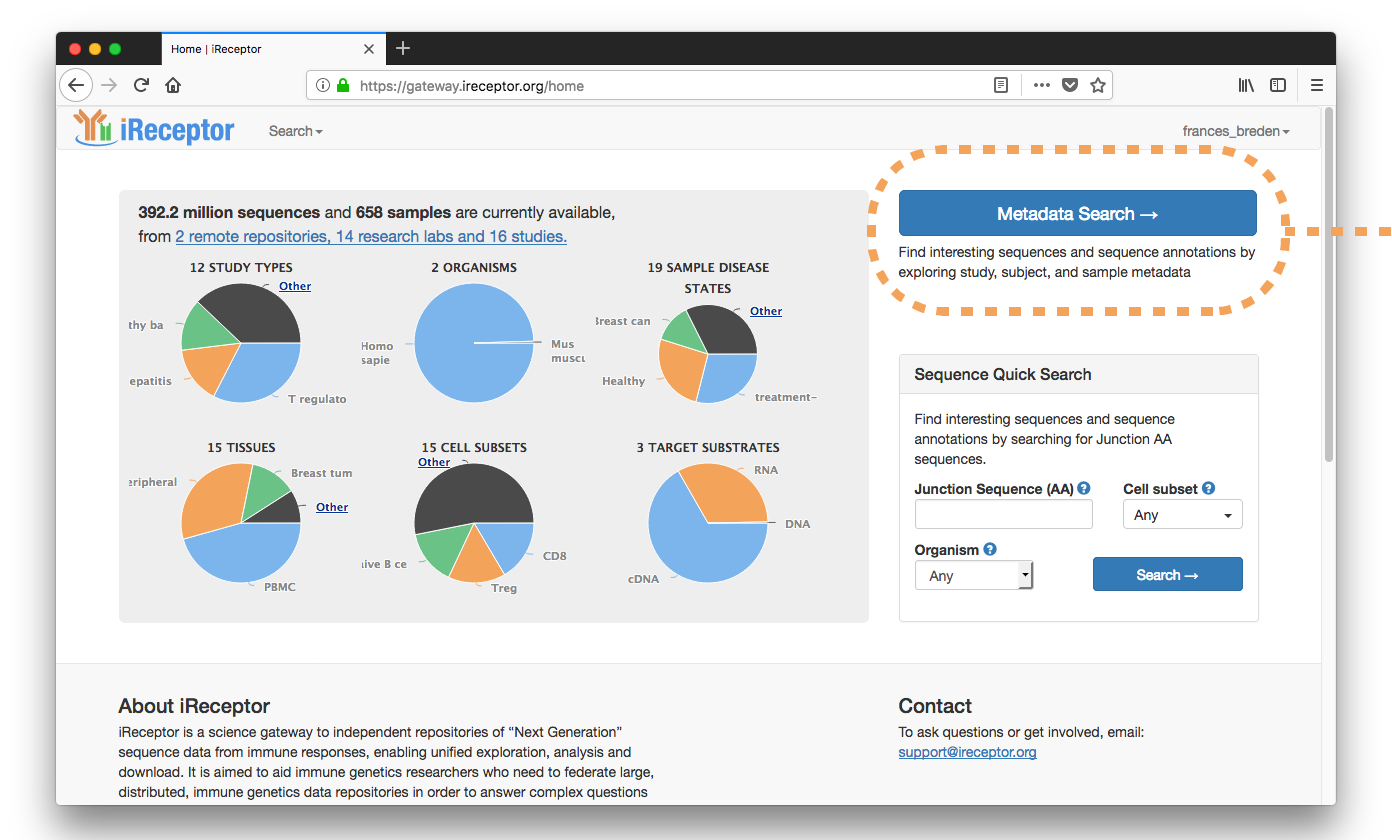
They enter the metadata search workflow and filter their results by Study Title ("child"), Study Type ("healthy"), and Cell Subset ("T Cell, Naive CD4+").
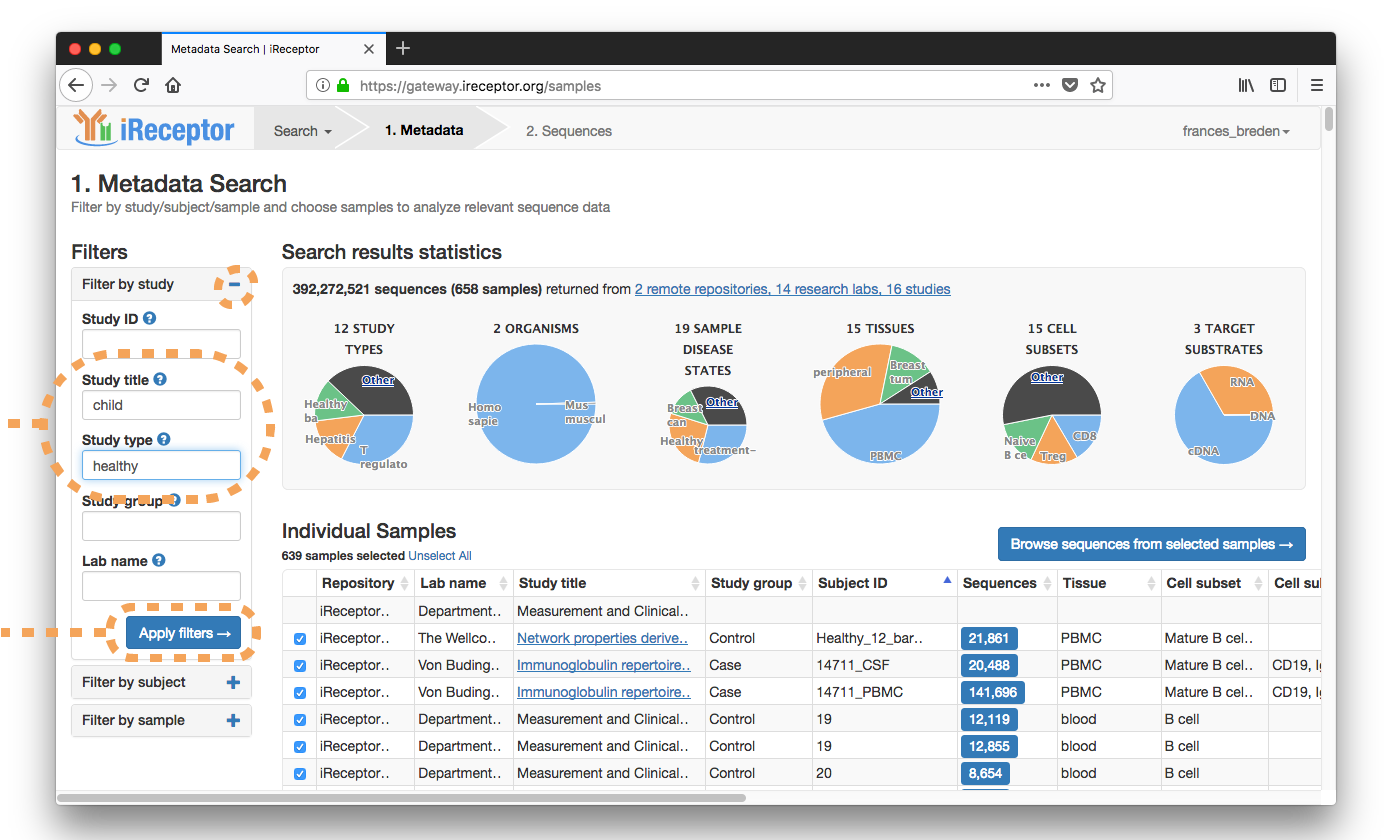
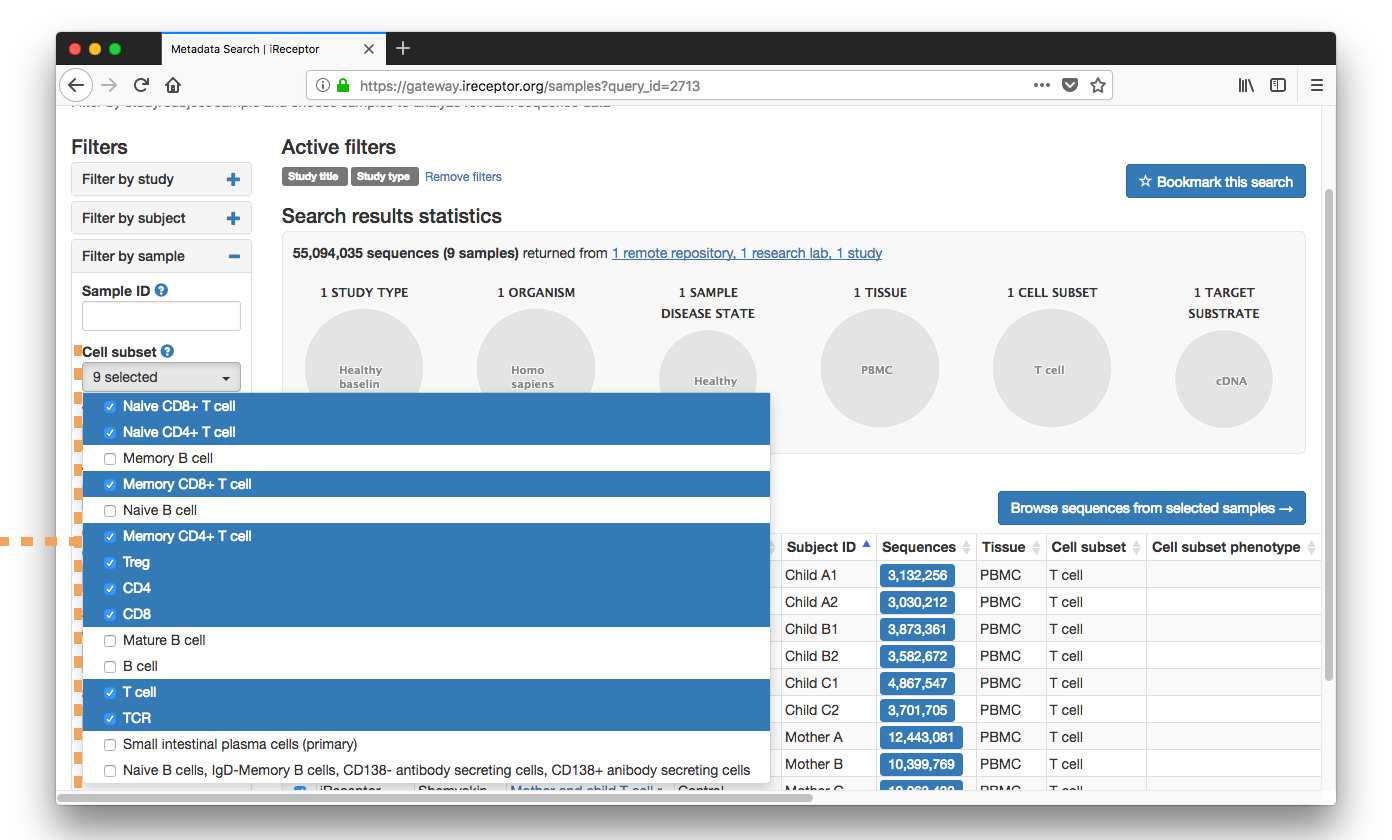
Results are returned from a paper on siblings' T cell repertoires from the iReceptor Public Archive in Vancouver, Canada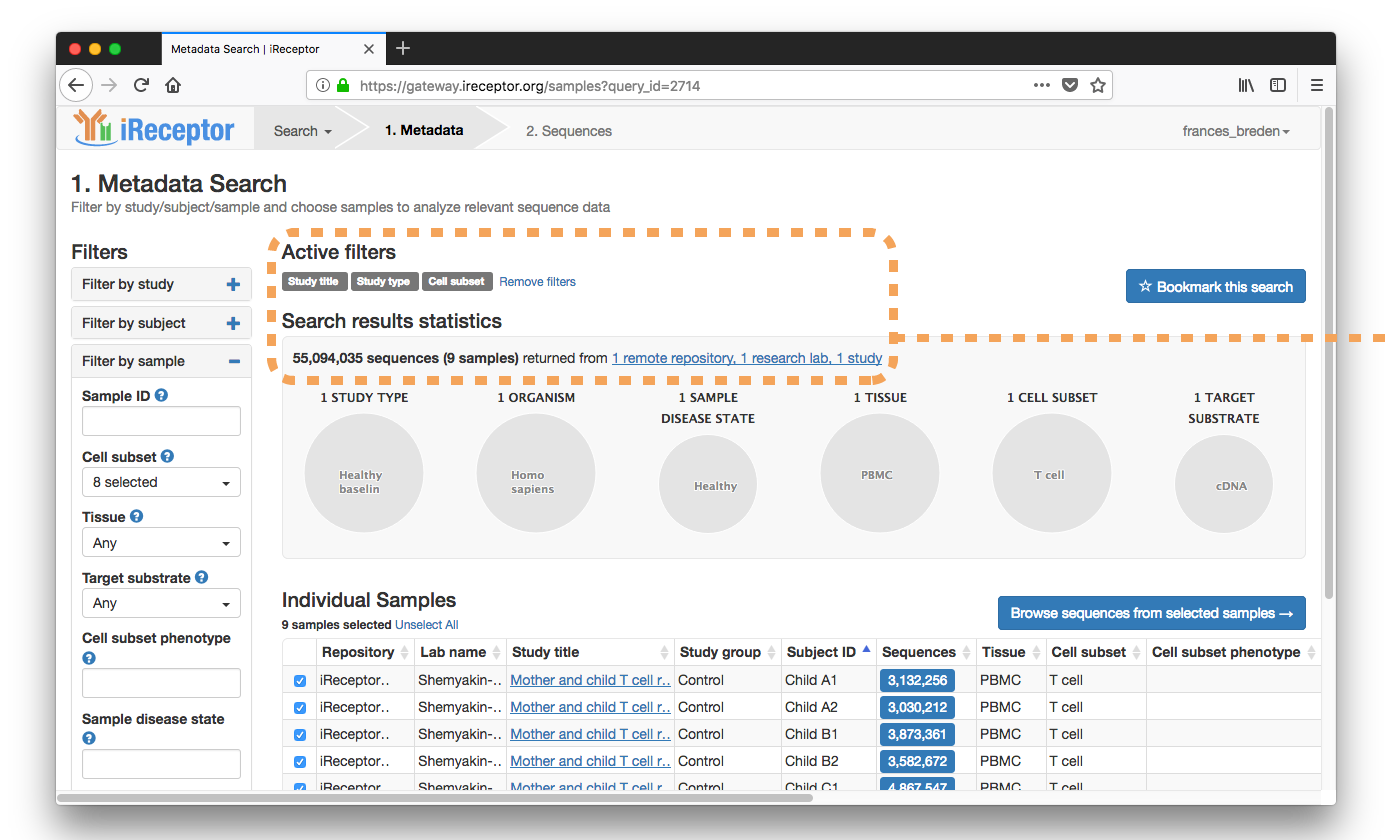
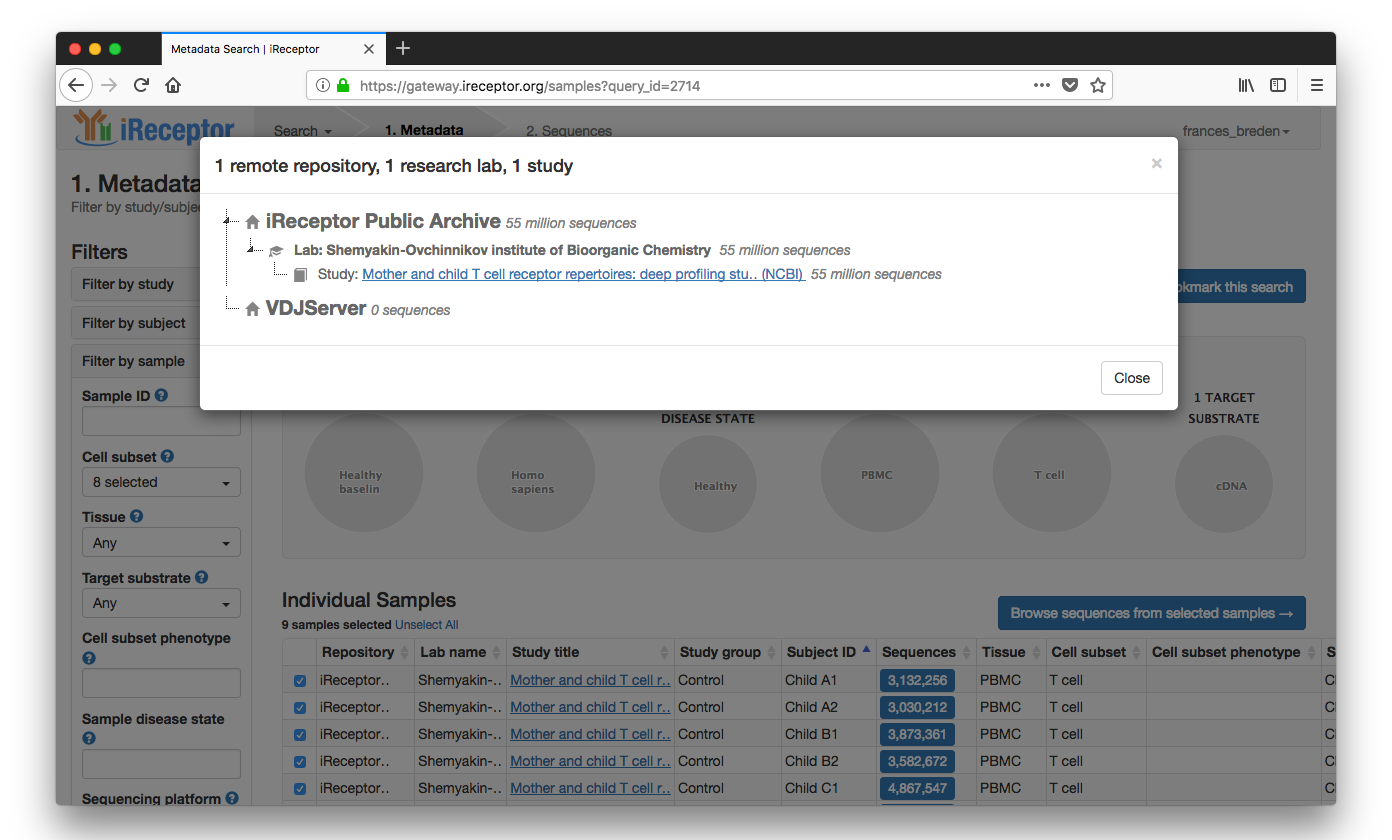
The researcher selects from the Individual Samples list sibling Sample IDs ("Child B1, Child B2")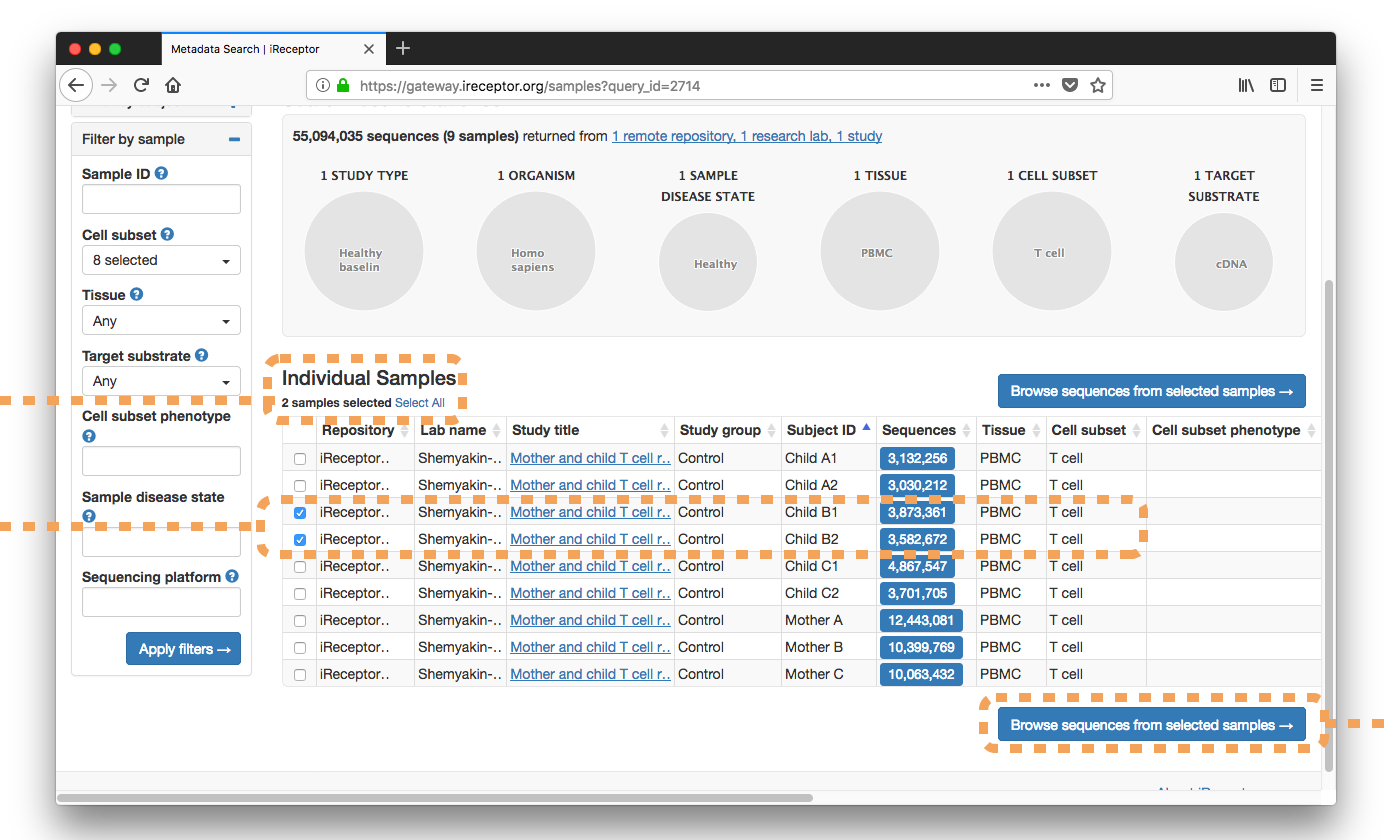
They download the resulting ~7,000,000 sequences for further analysis.
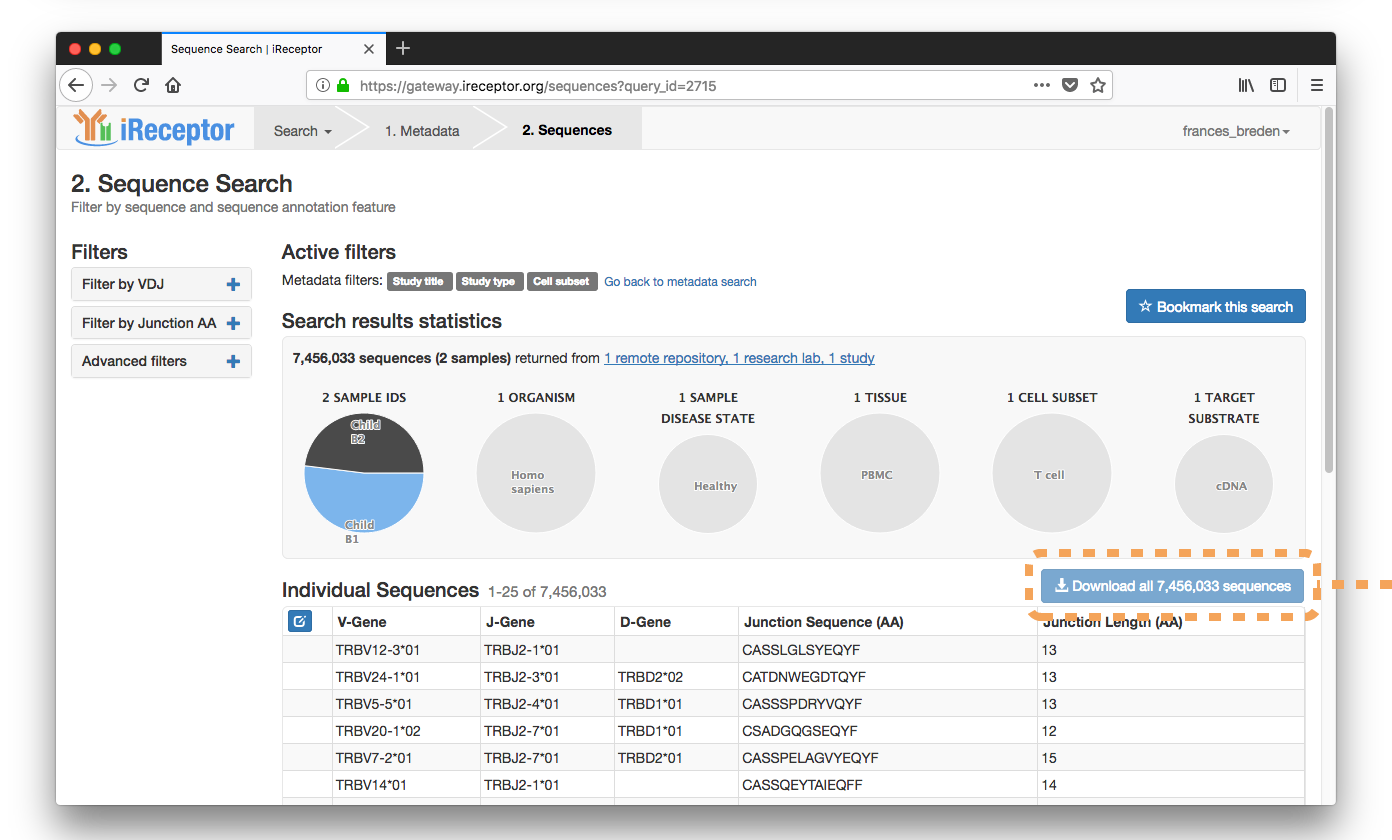
Use-case Scenario: Sequence Search
A reasearcher would like to know: is the following TCRB motif that is associated with Epstein Barr Virus (EPV) infection in lung translant recipients common in healthy patient populations?
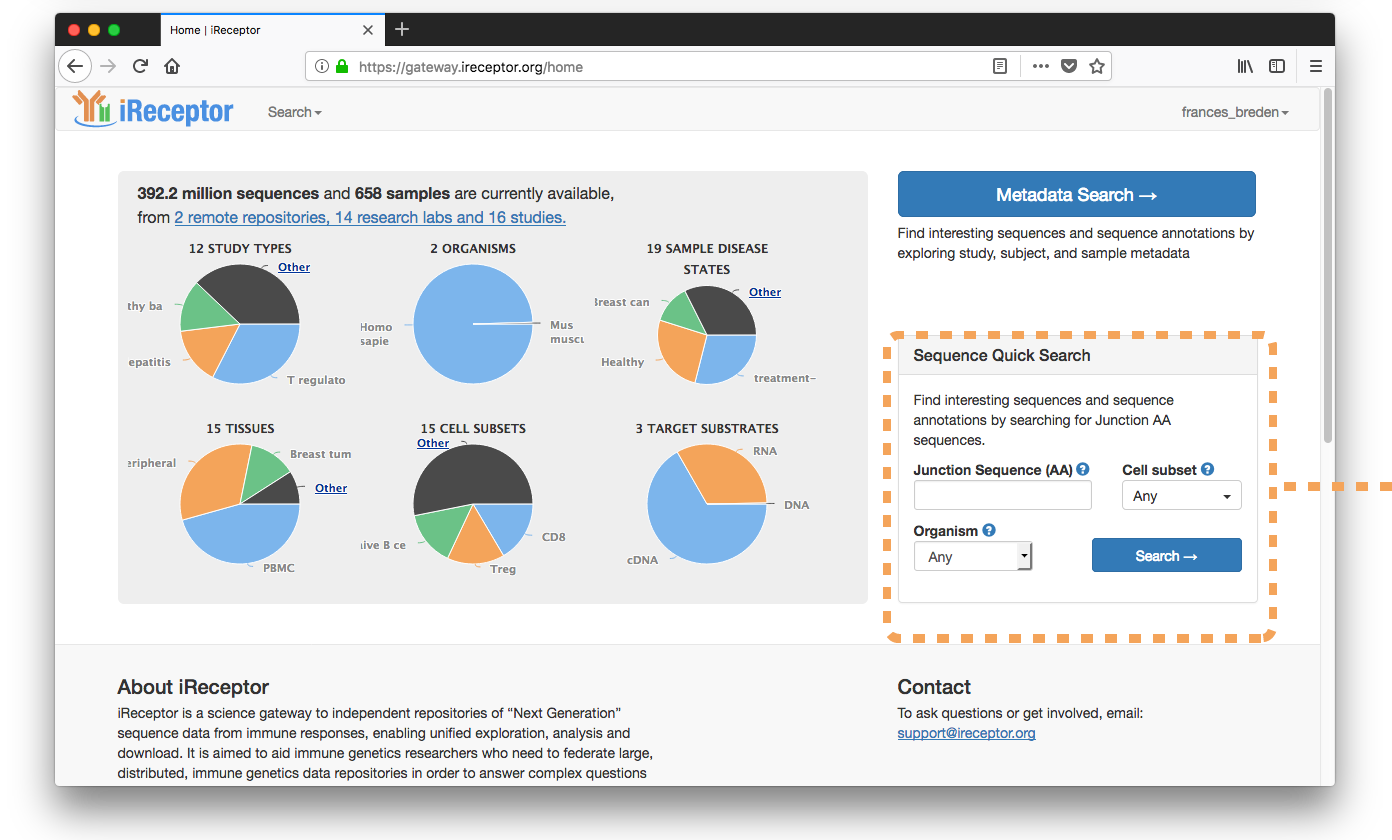
Searchers for EBV-associated Junction AA Sequence ("GTGGTNEKL")
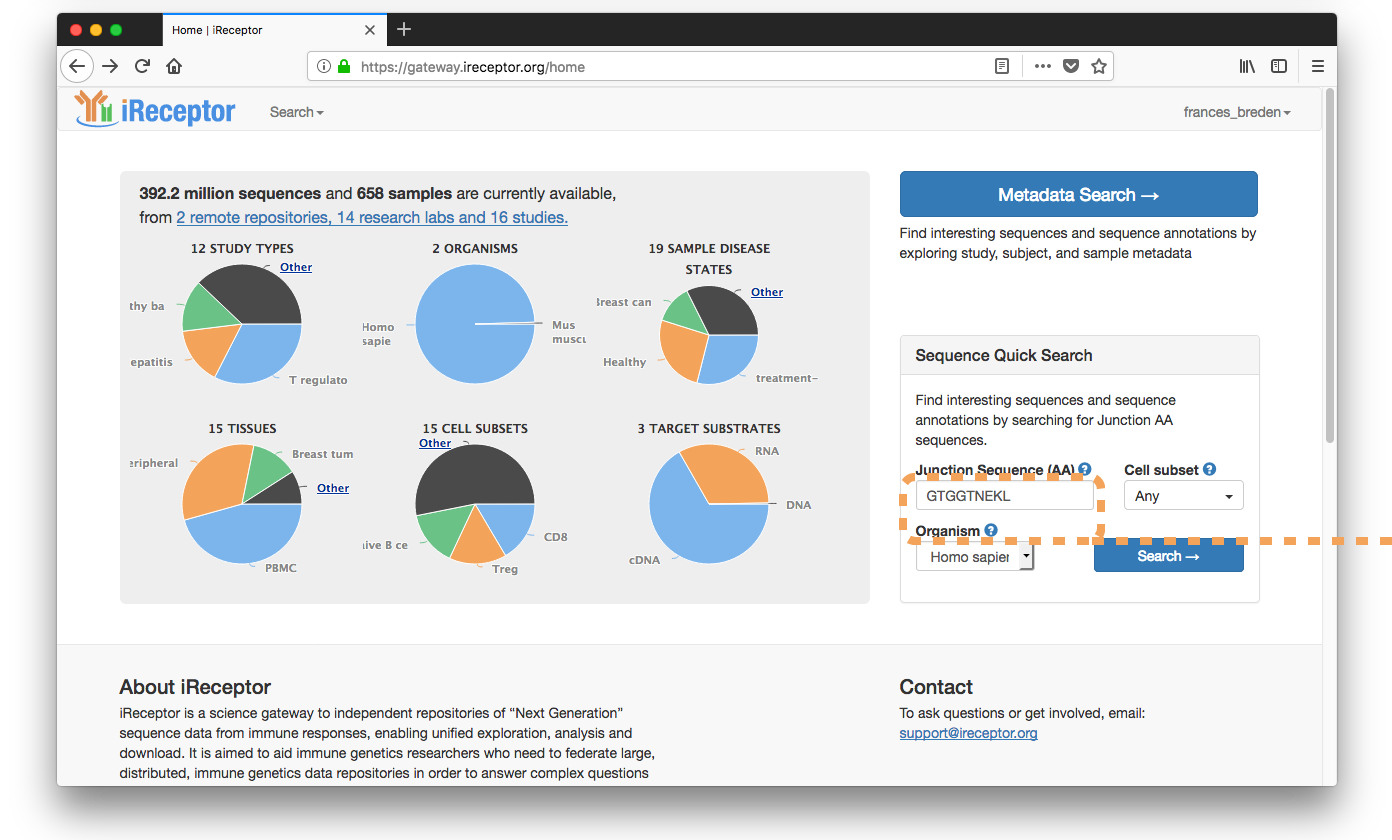
Discovers T cell repertoires from 8 healthy subjects with the sequence "GTGGTNEKL".
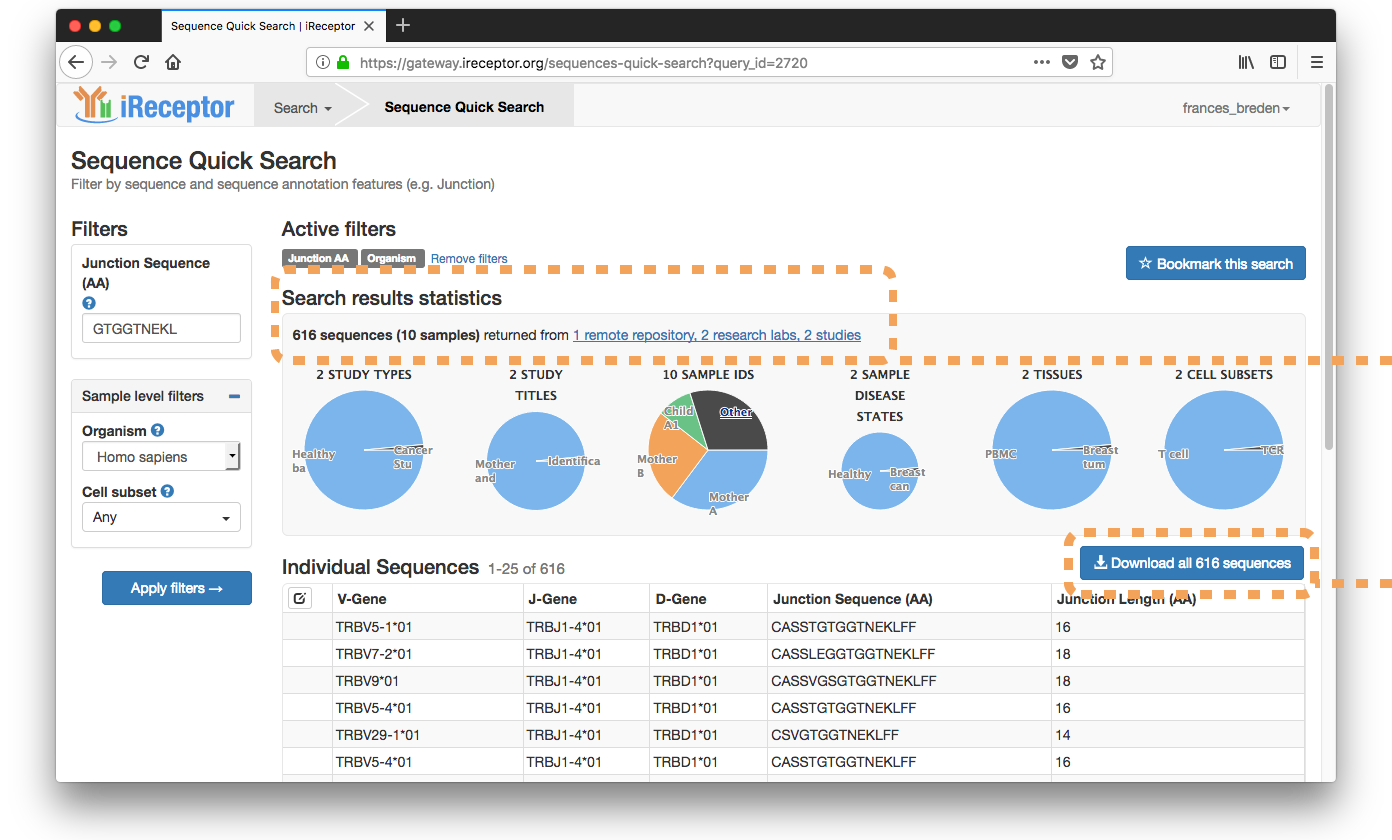
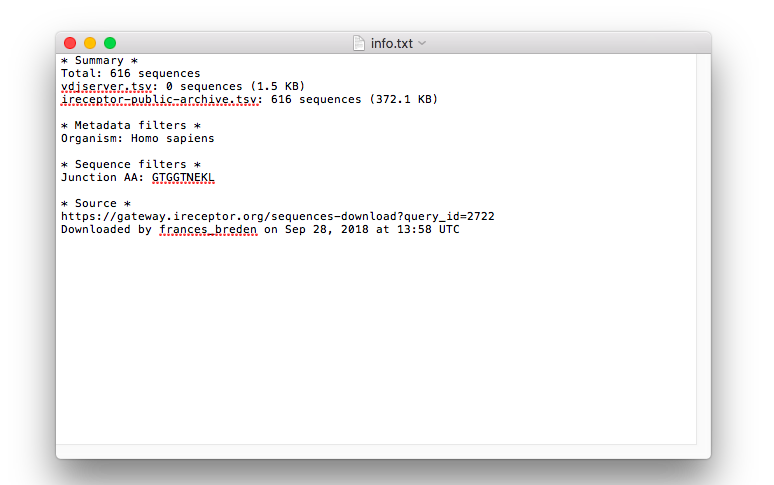
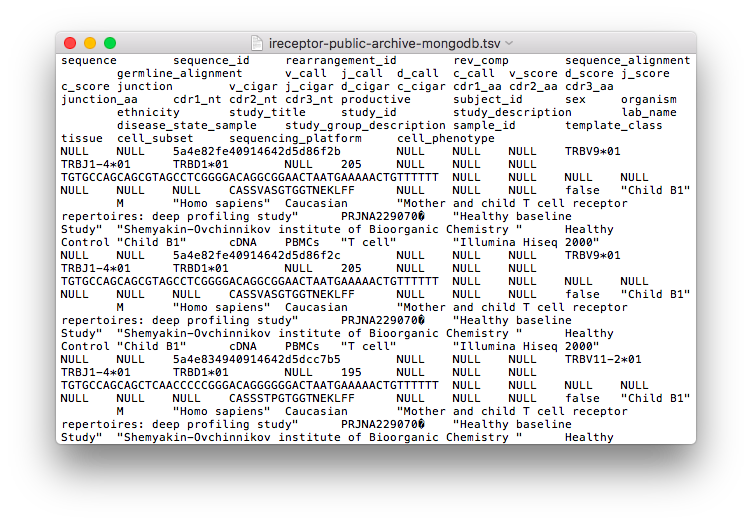
Downloads sequences for further analysis (pictured below: the readme information and a text version of the download results)