iReceptor is a data discovery platform that facilitates the curation, analysis and sharing of antibody/B-cell and T-cell receptor repertoires (Adaptive Immune Receptor Repertoire or AIRR-seq data) and related subject and sample metadata from multiple studies across many labs and institutions. We are committed to providing a platform for researchers to increase the value of their data through sharing with the community. This will greatly increase the amount of data available to answer complex questions about the adaptive immune response, accelerating the development of vaccines, therapeutic antibodies against autoimmune diseases, and cancer immunotherapies.
Curated Studies Available
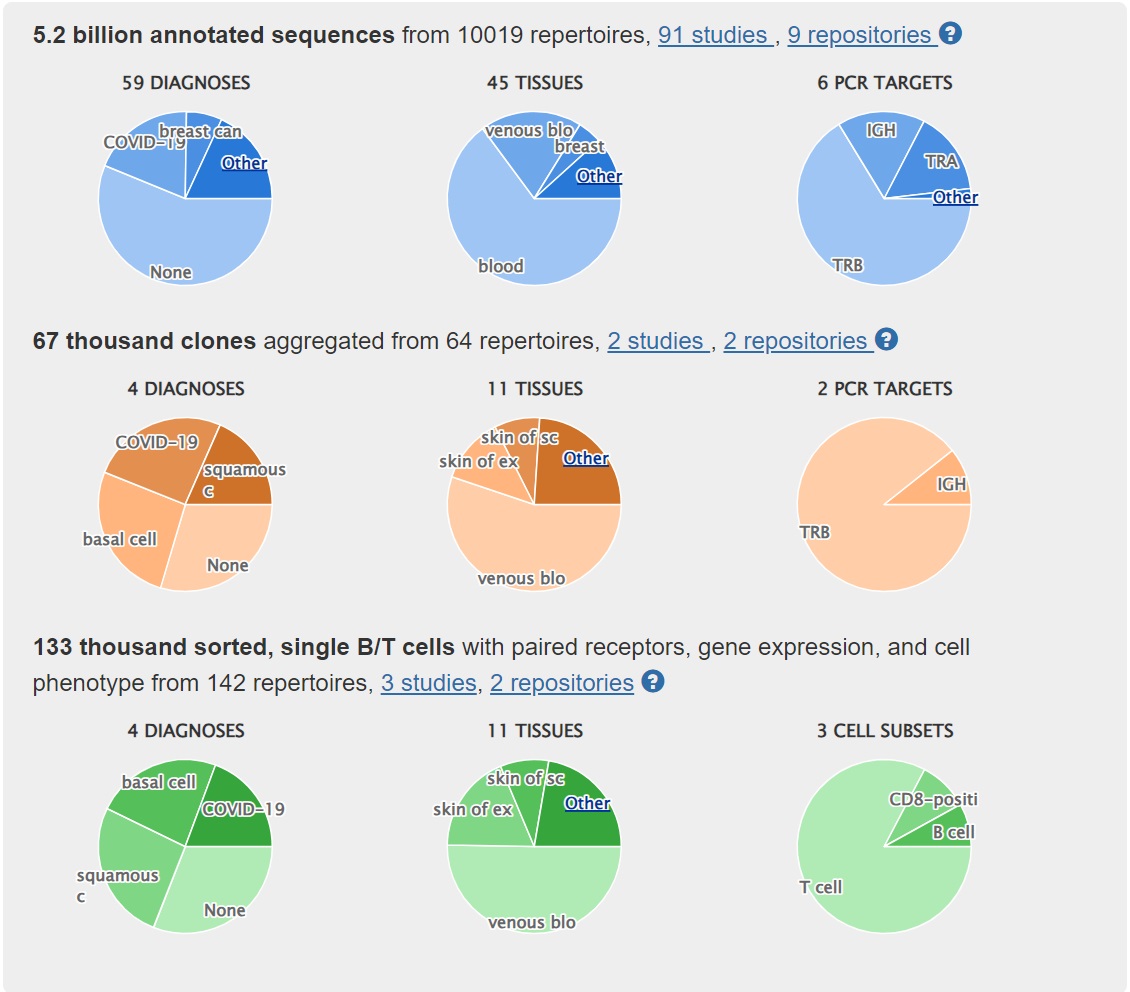
Users of the iReceptor Gateway can search AIRR-seq data from studies on cancer (for example breast and ovarian), autoimmune diseases (for example MS and SLE), and infectious diseases (for example HIV and influenza). Working with the COVID-19 and Type 1 Diabetes communities we have recently expanded our curated studies in those areas.
Following the AIRR Data Commons standards we are the only public resource that curates data from single-cell immune-profiling studies.
Visit the iReceptor Gateway to explore this data.
Our Approach
The iReceptor Gateway integrates large, distributed, AIRR-seq data repositories, following standards for sharing and interoperability developed by the Adaptive Immune Receptor Repertoire (AIRR) Community. The main goal of iReceptor is to connect this distributed network of AIRR-seq repositories into an AIRR Data Commons, allowing queries across multiple projects, labs, and institutions.
Present functionalities include:
- Search for repertoires satisfying certain metadata (e.g. find all AIRR-seq repertoires from ovarian cancer studies)
- Search for all repertoires that contain specific CDR3 sequences
- Search identified repertoires for sequences derived from particular V, D, and J genes and alleles
- Search identified repertoires for clones derived from particular V, D, and J genes and alleles
- Search identified repertoires for cells with specific cell and gene expression characteristics
- Download sequences, clones, or cells from these repertoires in AIRR formats, easily importable to other AIRR-seq analysis tools
- Directly perform analyses within the iReceptor Gateway on these sequences, cells, and clones
iReceptor is a member of the iReceptor Plus Consortium, resulting in the addition of Cell and Clone data as well as an integrated Analysis capbility, enbaling more powerful systems immunology approaches to AIRR-seq analysis.
Researchers interested in sharing data or exploring the AIRR Data Commons through the open iReceptor Gateway should visit www.ireceptor.org and for an account contact support@ireceptor.org. For more information about the iReceptor approach and architecture, please see the paper iReceptor: a platform for querying and analyzing antibody/B-cell and T-cell receptor repertoire data across federated repositories.
Funding
iReceptor is a project located at the IRMACS Centre at Simon Fraser University. It is funded by the CANARIE Network Enabled Platforms Program, the Canada Foundation for Innovation (CFI), the British Columbia Knowledge Development Fund (BCKDF), the Canadian Institute for Health Research, the European Union’s Horizon 2020 research and innovation programme (grant agreement No 825821), and Simon Fraser University. The iReceptor project was part of the EU/CIHR funded iReceptor Plus Consortium.
How to Acknowledge iReceptor
If you find that the iReceptor Gateway is useful in your research and/or you download and reuse data from the AIRR Data Commons, please cite the iReceptor and the AIRR Data Commons papers:
[Corrie et al.] iReceptor: a platform for querying and analyzing antibody/B-cell and T-cell receptor repertoire data across federated repositories, Immunological Reviews, Volume 284:24-41 (2018)
[Christley et al.] The ADC API: A Web API for the Programmatic Query of the AIRR Data Commons, Front. Big Data, 17 June 2020.
Suggested wording for a citation that is storing data in the AIRR Data Commons would be: "Study data is available for download from the AIRR Data Commons [Christley et al.] using the iReceptor Gateway [Corrie et al.] using Study ID PRJNA00000.”
Suggested wording for an attribution for data reuse would be "Study data was queried and downloaded from the AIRR Data Commons [Christley et al.] using the iReceptor Gateway [Corrie et al.] on January 1, 2021.”