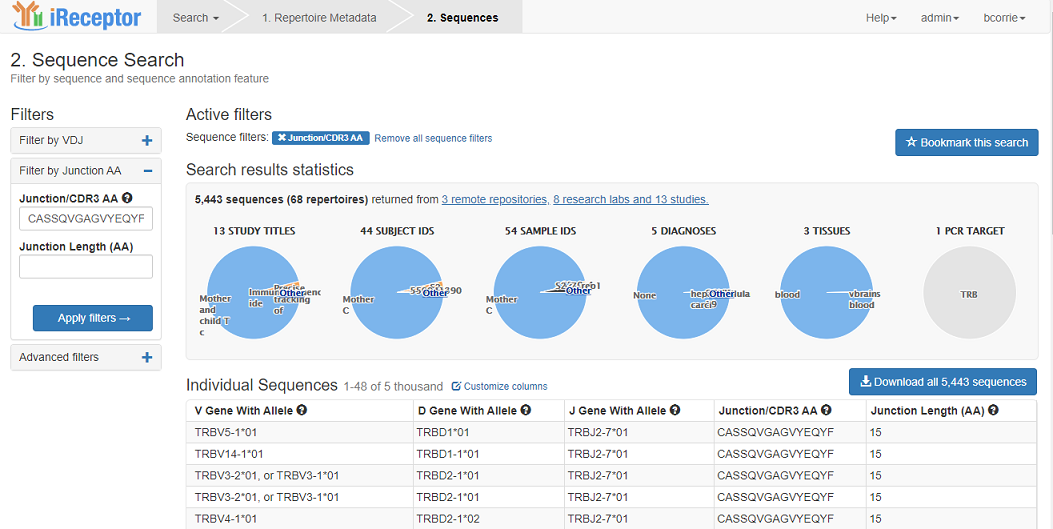
At the end of your sequence rearrangement search, you can download the results of your search and the associated metadata using the download button.
Downloads page
This will take you to the download page, which lists the set of downloads that you have recently performed. You can return to the download page at any time through the User menu in the top right corner of the iReceptor Gateway page.
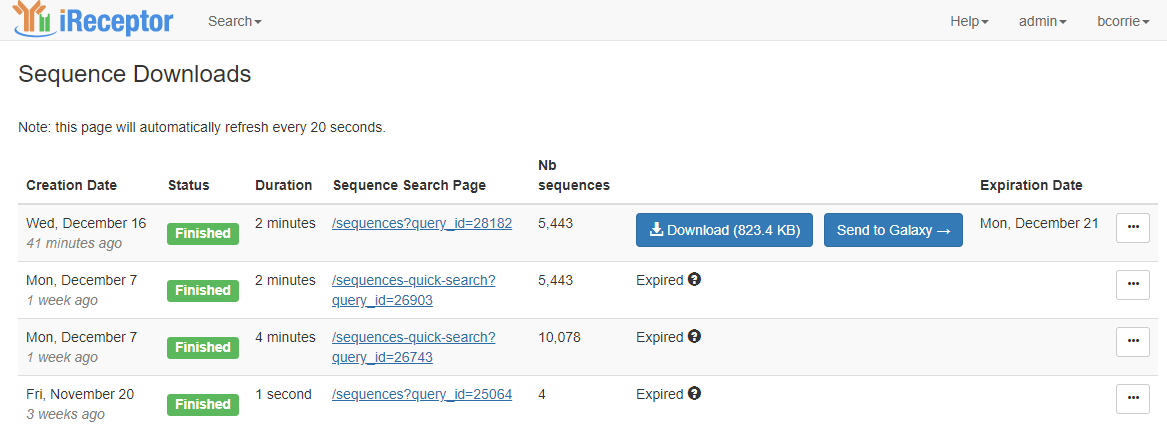
Downloads are an asynchronous process and are queued to run in order of submission. When your download is queued it will be in the "Queued" state and it will show you how many downloads are in the queue in front of you. Once your download starts it will be in the "Running" state. This means that the iReceptor Gateway is searching the repositories and federating the data into a single download bundle. This may take some time depending on the size of your download. Once the download to the Gateway is completed, the download will be in the "Finished" state, and you will then be able to click on the "Download" button. You will receive an email when the iReceptor Gateway is done federating your data. When you click the "Download" button the data will then be downloaded from the iReceptor Gateway to your local computer. The "Download" button will tell you how large the download is. The iReceptor Gateway keeps downloads available for a fixed period of time (currently approximately a week) and then removes them, so you need to download the data promptly. The download page does keep the query used to generate the download, so you can revisit the page and redo the data federation if need be.
Download format
You will receive a ZIP archive with the results of your download request. The ZIP archive will consist of a single "info.txt" file that will provide a summary of your download request. In addition, for each repository that contained data that met your query constraints you will receive an AIRR TSV rearrangement file and an AIRR Repertoire JSON file. For example, if your download request found data in three repositories, you would receive 3 AIRR TSV files, 3 AIRR Repertoire JSON files, and one "info.txt" file.
The info.txt file will look something like the following, providing a summary of what was downloaded, which filters were used, and a link to the user query that generated the download (for reproducibility).
* Summary * Total: 18252 sequences airr-covid-19-1.tsv: 18252 sequences (6.4 MB) * Metadata filters * short.sort_column: ir_sequence_count short.sort_order: asc short.cols: rest_service_name,study_title,disease_diagnosis,study_group_descript ion,ir_sequence_count,lab_name,tissue,pcr_target_locus,cell_subset,cell_phenotyp e,pub_ids,study_id,subject_id,sample_id,template_class,sequencing_platform * Sequence filters * None * Source * /sequences?query_id=XXXX Downloaded by XXXX on Jul 22, 2020 at 16:08 UTC
The files that you might expect from a download from a single repository might look something like this:
user:~$ ls airr-covid-19-1-metadata.json airr-covid-19-1.tsv info.txt ir_2020-07-22_1608_5f18647f4ef8f.zip
The AIRR Repertoire JSON file from each repository will contain a JSON object for each Repertoire. This JSON object will contain all of the Repertoire metadata for that Repertoire. Each Repertoire object contains three identifier fields (repertoire_id, data_processing_id, and sample_processing_id) that link a Repertoire object to the Rearrangements in the AIRR TSV rearrangement file. Each row in the AIRR TSV files contains rearrangement data as described in the AIRR Communities AIRR TSV specification. Each rearrangement has at least one of the three identifier fields and is guaranteed to always have a repertoire_id. This enables you to link a specific Rearrangement with a specific Repertoire and will allow you to process the data based on Repertoire if required. A specific Repertoire can contain biological samples that are processed differently (identified by sample_processing_id) or sequence data that is annotated differently (identified by data_processing_id) so all three identifiers may be required to link a specific rearrangement record in the AIRR TSV file to a specific set of Repertoire metadata.
Processing AIRR download data
One common workflow for processing data that is downloaded via the iReceptor Gateway would be to download some data of interest, split that data based on repertoire_id into separate files, and then perform some comparative analysis between the Repertoires of interest. For example, if you wanted to identify a set of Rearrangements for a specific Repertoire you would need to identify the Repertoire of interest in the Repertoire JSON file, find its repertoire_id, and then filter the Rearrangements in the AIRR TSV file based on that repertoire_id. If you wanted to specifically focus on one subset of the data based on Sample Processing protocols (via sample_processing_id) or Data Processing protocols (via data_processing_id), you would need to filter the data on more than one of the identifier fields.
